AI that sheds light on living organisms: at CBIO, deciphering biology to improve healthcare

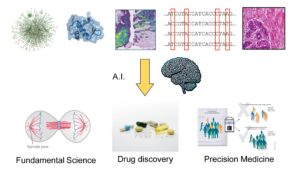
The research conducted at CBIO lies at the intersection of mathematics, computer science, biology, and medicine. Its mission is clear: to develop machine learning and AI methods capable of making sense of contemporary biological data, which is growing in both quantity and complexity.
This work addresses three major scientific objectives:
To achieve these objectives, the CBIO relies on a team of permanent researchers with complementary expertise: Chloé-Agathe Azencott, Éloïse Berson, Florian Massip, Vincent Mallet, Véronique Stoven, and Thomas Walter. The CBIO has a strategic partnership with the Institut Curie, a research center and hospital dedicated to cancer, and is affiliated with the “Computational Oncology” unit, which enables it to tackle complex problems and respond to specific clinical needs in oncology.
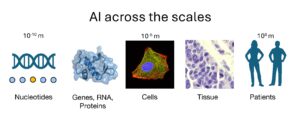
Being able to predict the progression of a disease or the effectiveness of a treatment is now a major challenge in oncology. AI offers powerful methods for integrating multiple data sources and producing such predictions. However, a key obstacle to these approaches lies in the very nature of the data being analyzed. We refer to data as very high-dimensional when a very large number of variables are measured for a limited number of patients or samples.
For example, in genomic studies, the objective is to identify statistical links between mutations—often numbering in the millions—and a clinical phenotype. In transcriptomics, the expression of approximately 20,000 genes is analyzed. In both cases, the number of patients from which AI models can learn is typically several orders of magnitude smaller than the number of variables measured, making traditional analysis methods fragile and sometimes misleading.
The CBIO therefore adopts several complementary strategies to address this problem:
These approaches not only enable prediction, but above all identify the biological mechanisms actually involved, which is a key issue for biomedical research.
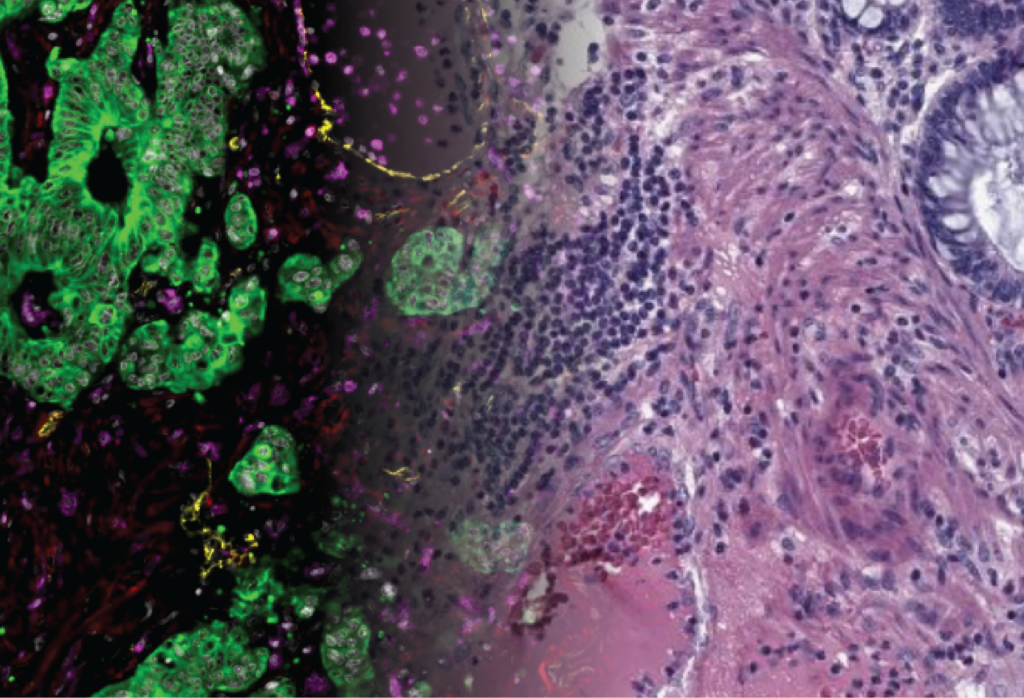
One of CBIO’s most visible areas of research is computational pathology, which uses AI to analyze images of biological tissue from histopathology, a discipline that uses microscopic examination of tissue for diagnostic purposes. These images, obtained from stained tissue sections and observed under a microscope, are now digitized in the form of whole-slide images that can reach several gigabytes in size.
The challenge is to automatically extract molecular or clinical information from tissue morphology alone.
CBIO researchers have developed methods capable of:
In concrete terms, the images are divided into thousands of small regions, analyzed by AI models pre-trained on millions of medical images, and then aggregated to produce a patient-level prediction. This work paves the way for faster, less expensive, and more accessible oncology, without the need for systematic, labor-intensive molecular analyses. In all of this work, the CBIO collaborates closely with hospital pathology departments, notably at the Institut Curie with Anne Vincent Salomon, Director of the IHU Cancers des Femmes (Institut Curie – Université PSL – Inserm) and chief pathologist at the Pôle de Médecine diagnostique et théranostique (PMDT), as well as Yves Allory, head of the “Molecular Oncology” research team within the Cell Biology and Cancer Unit (UMR144) and head of the Pathology Department.
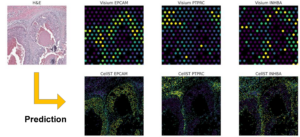
Another major advance concerns spatial transcriptomics, a recent technology that makes it possible to observe not only which genes are expressed in a tissue, but also where exactly they are expressed. In concrete terms, this approach associates each point in a tissue with a measurement of the activity of thousands of genes, sometimes up to 20,000 genes at a time, while retaining their exact position in space. It thus offers a detailed view of the architecture of a tissue, linking the structure of the tissue to its molecular functioning.
These data are particularly rich, but they are also complex and costly to produce and analyze. It raises new computational challenges, such as cellular deconvolution, which involves determining which cells contribute to a signal measured in a given area, or the integration of several types of data, such as tissue images and molecular measurements. Another challenge is to establish predictive links between different modalities in order to supplement one piece of information with another.
To address these challenges, researchers at CBIO and the Systems Biology team at the Institut Curie, led by Emmanuel Barillot, have developed AI models capable of predicting gene expression at the cellular level based on the visual appearance of tissue observed under a microscope. These models can estimate highly detailed molecular information from routine clinical examinations, without systematically resorting to heavy and costly technologies.
These approaches open up prospects for integrative spatial biology combining images, genetic data, and clinical information. They are particularly promising for better understanding complex diseases such as cancer, where the spatial organization of cells plays a key role in disease progression and response to treatment.

CBIO research also covers structural biology, which focuses on the three-dimensional shape of proteins and RNAs. Thanks to AI, it is now possible to:
At the same time, the center is developing methods to reconcile AI statistical models with mechanistic models of biology, in order to move from correlation to understanding of the underlying biological mechanisms.
This work was highlighted during the AI Workshop held in December 2025 at Mines Paris – PSL. Designed as an opportunity for internal exchange, the event allowed faculty, doctoral students, and engineers to present their projects, tools, and platforms through oral presentations and posters. Beyond the diversity of topics, the workshop highlighted a common dynamic: building AI rooted in reality, integrating it into complex systems, and tackling major societal challenges.
By developing robust, explainable AI rooted in biological reality, CBIO is helping to transform the way life sciences are studied and applied. From fundamental research to the discovery of treatments and precision medicine, its work illustrates how AI can become a tool for understanding living organisms, serving concrete and responsible medical innovations.

Understanding, predicting, and optimizing the behavior of materials is a key challenge for industry, from aeronautics to energy, transportation, and a...